VRprofile2: detection of antibiotic resistance-associated mobilome in bacterial pathogens
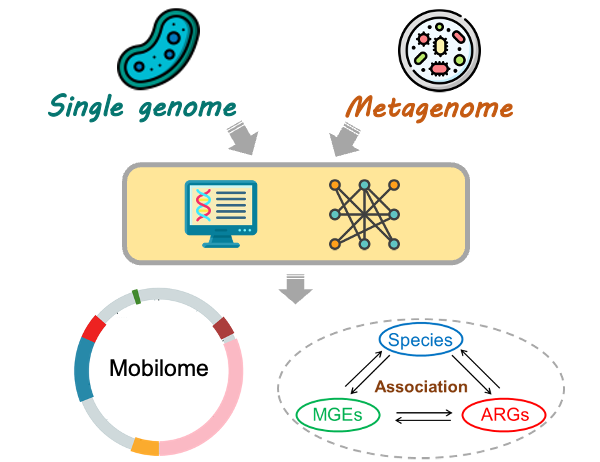
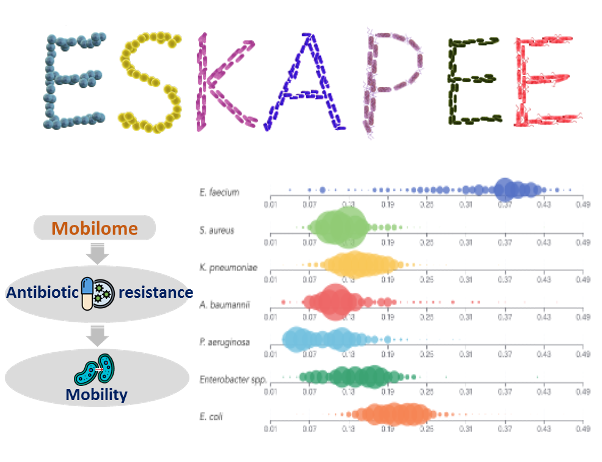
Antibiotic resistance has become one of the biggest threats to public health globally. The antibiotic resistance genes (ARGs) often gain mobility through associating with a set of mobile genetic elements (mobilome). Their concerted activities significantly contribute to the promiscuous spread of ARGs throughout the microbial community. Here, we report the release of VRprofile version 2.0, which offers three major enhancements: (i) graphical representation of the multiple resistance regions with mosaic structures and comparison to the known MGEs with similar architecture; (ii) aid to explore the relationships of antibiotic resistance genes, mobile elements, and host strains; (iii) pre-computed mobilome of more than 5500 ESKAPEE bacterial genomes. We expect that VRprofile2 will provide better support for researchers interested in bacterial diverse mobile elements and the dissemination of antibiotic resistance.